Chapter 4 Exercise 9.5
4.1 Load packages
Here is the R code to download the required packages for this exercise.
## Loading required package: pacman4.2 Data
This is equivalent to data step in SAS. Here, the data is imported from a file data.csv using the function read_csv. This function will download the file directly from here.
# Import data
a <- read_csv("https://raw.githubusercontent.com/luckymehra/epidem-exercises/master/data/ex9_5.csv")## Parsed with column specification:
## cols(
## COL = col_double(),
## ROW = col_double(),
## YI = col_double()
## )## # A tibble: 144 x 3
## COL ROW YI
## <dbl> <dbl> <dbl>
## 1 1 1 2
## 2 2 1 2
## 3 3 1 0
## 4 4 1 3
## 5 5 1 1
## 6 6 1 1
## 7 7 1 1
## 8 8 1 5
## 9 9 1 22
## 10 10 1 13
## # ... with 134 more rows4.3 Autocorrelation statistics
# visualize the data
ggplot(data = a) +
geom_point(mapping = aes(x = COL, y = ROW, size = YI, color = YI)) +
ggtitle("Spatial Distribution of YI Observation") +
theme(plot.title = element_text(hjust = 0.5))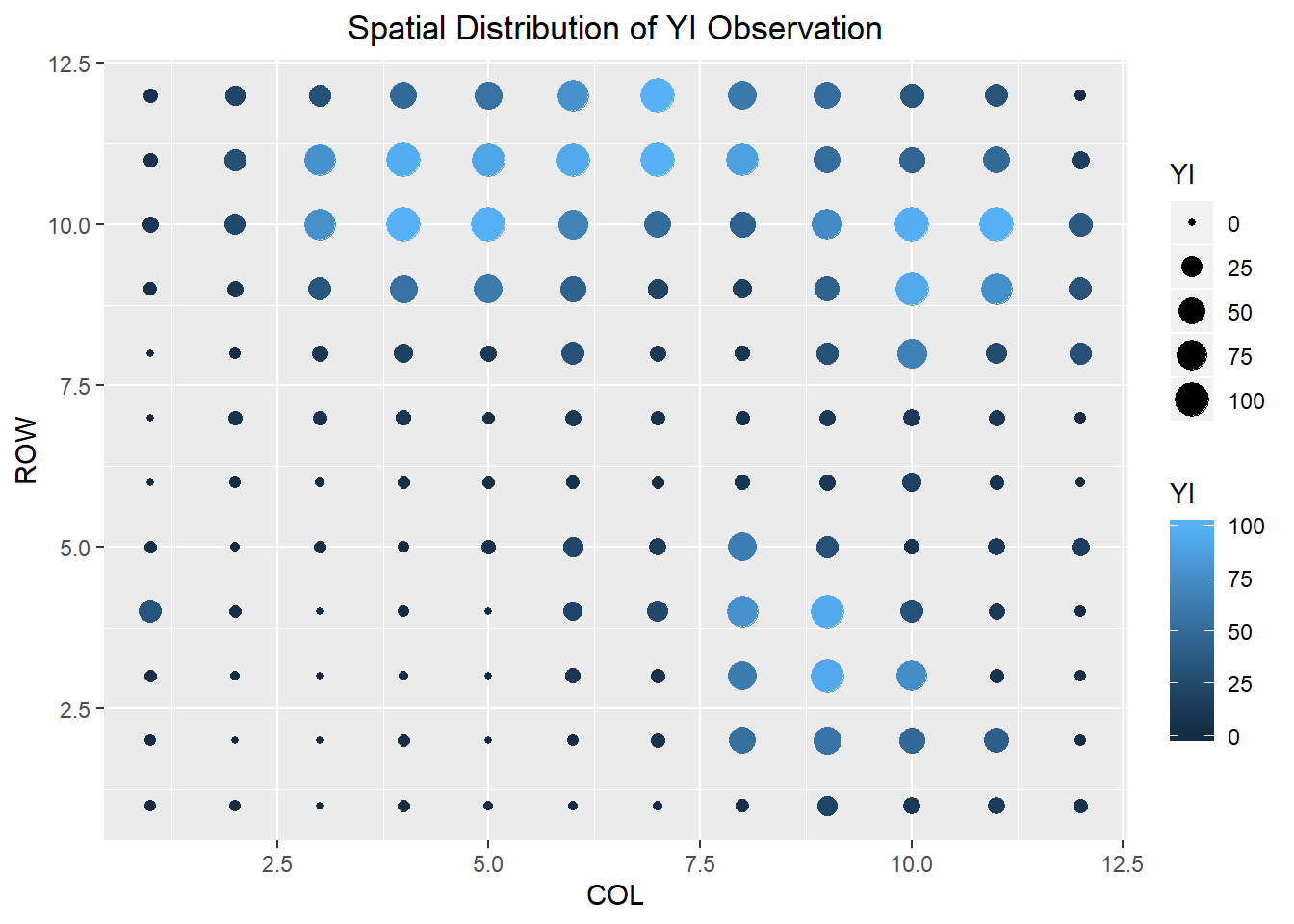
# calculate Moran's I
Coords <- a %>%
dplyr::select(COL, ROW)
mI <- moransI(Coords, Bandwidth = 1, a$YI)
# print Moran's I table
moran.table <- tribble(
~`Moran's I`, ~`Expected I`, ~`Z randomization`, ~`P value randomization`,
#------------/--------------/-------------------/------------------------
mI$Morans.I, mI$Expected.I, mI$z.randomization, mI$p.value.randomization
)
moran.table## # A tibble: 1 x 4
## `Moran's I` `Expected I` `Z randomization` `P value randomization`
## <dbl> <dbl> <dbl> <dbl>
## 1 0.782 -0.00699 13.0 1.28e-38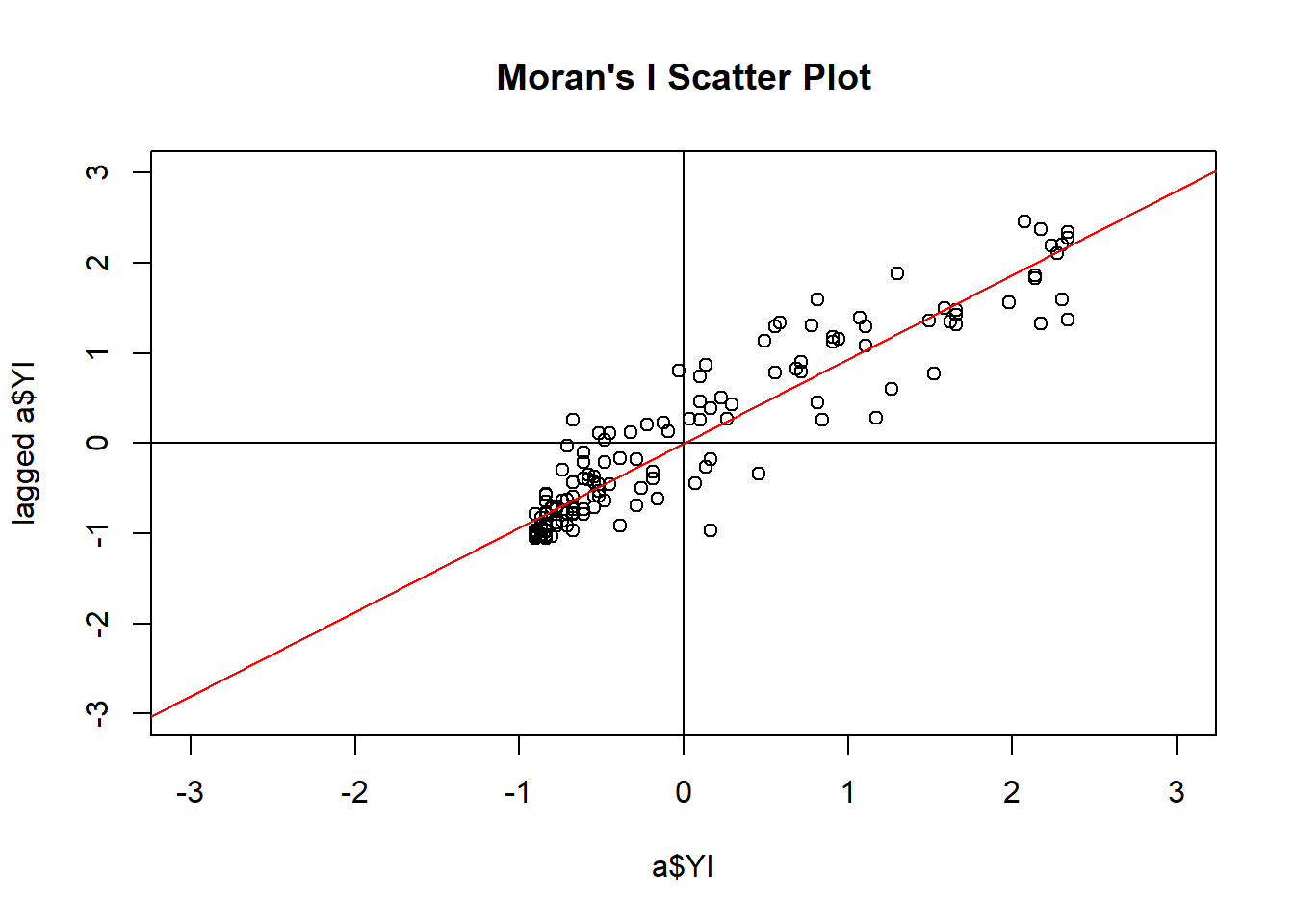
# calculate geary's c
Coords_num <- coordinates(Coords)
# create an object of class 'nb' so that it can be used with function from packege `spdep`
Coords_nb <- knn2nb(knearneigh(Coords_num))
# create a 'listw' object for use in the function `geary.test`
coords_listw <- nb2listw(Coords_nb)
gearyC <- geary.test(a$YI, coords_listw, alternative = "two.sided")
gearyC##
## Geary C test under randomisation
##
## data: a$YI
## weights: coords_listw
##
## Geary C statistic standard deviate = 8.8657, p-value < 2.2e-16
## alternative hypothesis: two.sided
## sample estimates:
## Geary C statistic Expectation Variance
## 0.235058006 1.000000000 0.0074444574.4 First variogram
We will use the package geoR to construct empricial variogram, and then draw them using package ggplot2.
## variog: computing omnidirectional variogram# extract data from object v1 for plotting
v1_plot_data <- cbind(v1$u, v1$v, v1$n) %>%
as.data.frame() %>%
dplyr::rename(Distance = V1,
Semivariance = V2,
Pair_count = V3)
# in the table below, gamma is semivariance
v1_plot_data## Distance Semivariance Pair_count
## 1 1 311.7154 506
## 2 2 636.2074 680
## 3 3 818.7044 812
## 4 4 901.3218 1386
## 5 5 910.2773 1044
## 6 6 942.3219 1252
## 7 7 998.2290 1046
## 8 8 1131.2105 1012
## 9 9 1261.6817 1076
## 10 10 1219.6067 614
## 11 11 1166.5541 508
## 12 12 910.6250 80# plot variogram
v1_plot_vario <- ggplot(data = v1_plot_data) +
geom_point(mapping = aes(x = Distance, y = Semivariance)) +
ggtitle("Empirical Semivariogram of YI") +
theme(plot.title = element_text(hjust = 0.5))
# plot pair counts
v1_plot_pair_count <- ggplot(data = v1_plot_data) +
geom_col(mapping = aes(x = Distance, y = Pair_count), width = 0.01, color = "blue")
# stack two plots
grid.arrange(v1_plot_vario, v1_plot_pair_count,
ncol = 1, heights = c(3, 1))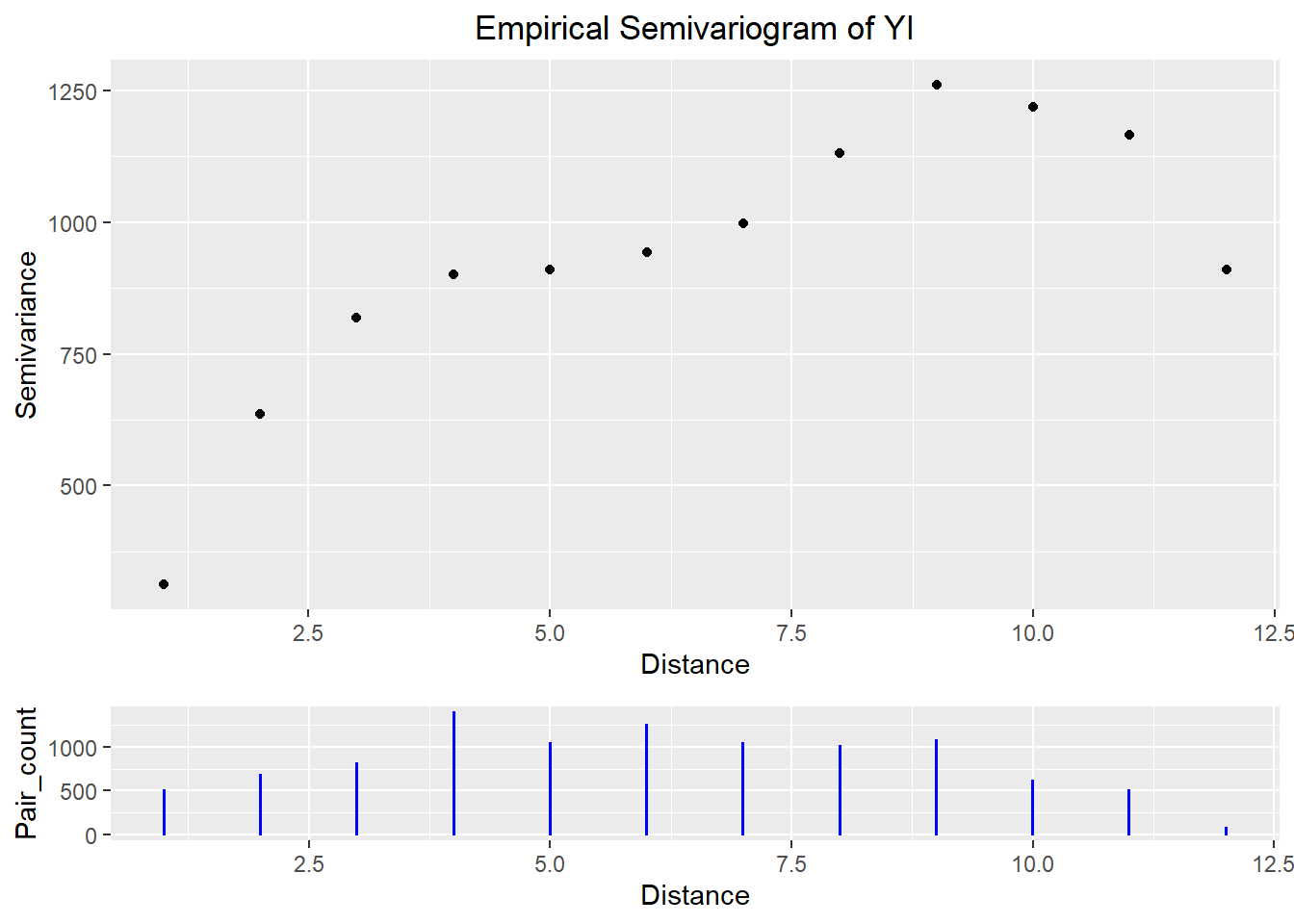
4.5 Second variogram
Plot robust and classical variogram together.
# fit robust variogram
v1_robust <- variog(coords = Coords_num, data = a$YI, breaks = seq(0.5, 15.5),
max.dist = 12, estimator.type = "modulus")## variog: computing omnidirectional variogram# extract the data
v1_robust_data <- cbind(v1_robust$u, v1_robust$v, v1_robust$n) %>%
as.data.frame() %>%
dplyr::rename(Distance = V1,
Semivariance = V2,
Pair_count = V3)
# plot robust variogram
v1_robust_vario <- ggplot(data = v1_robust_data) +
geom_point(mapping = aes(x = Distance, y = Semivariance)) +
ggtitle("Empirical Semivariogram of YI - Robust estimation") +
theme(plot.title = element_text(hjust = 0.5))
v1_robust_vario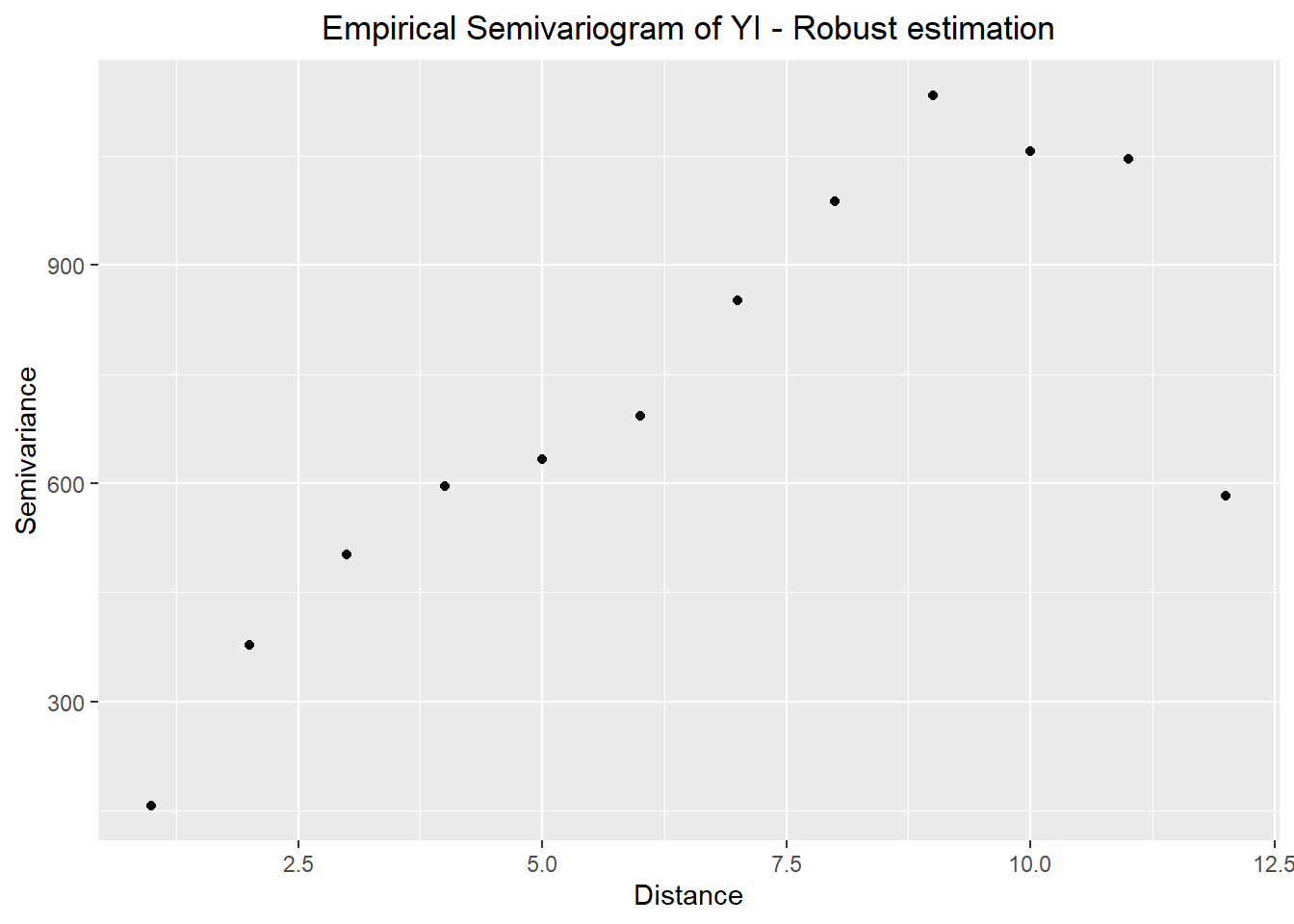
# combine robust and classical variogram
var_comb <- v1_robust_data %>%
# combine robust and classical variogram datasets
dplyr::rename(Semivariance_robust = Semivariance) %>%
bind_cols(dplyr::select(v1_plot_data, Semivariance)) %>%
gather(key = "Semivariance_type", value = "Semivariance", -c(Distance, Pair_count)) %>%
# plot
ggplot() +
geom_point(mapping = aes(x = Distance, y = Semivariance, color = Semivariance_type)) +
ggtitle("Empirical Semivariogram for YI") +
theme(plot.title = element_text(hjust = 0.5))
var_comb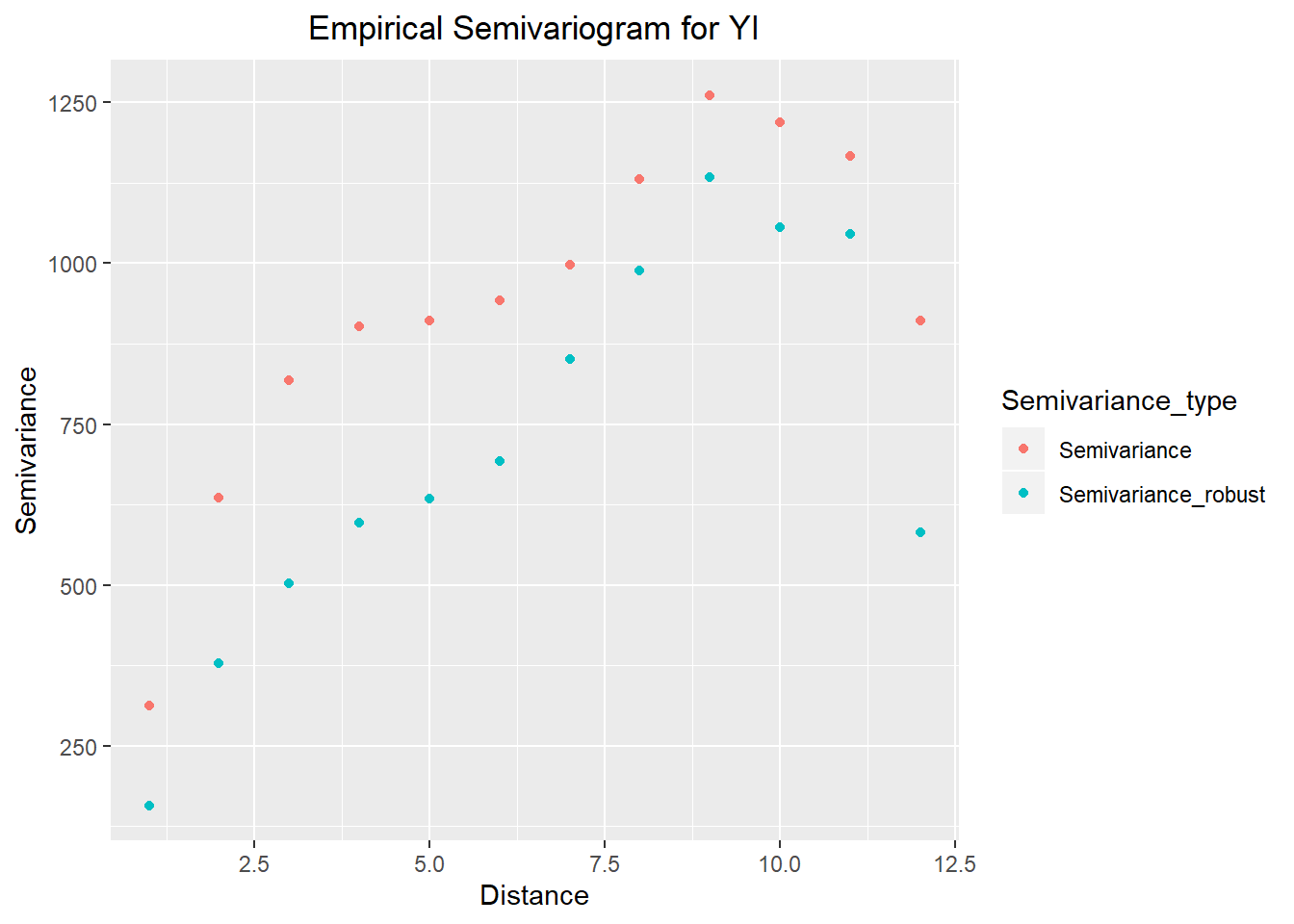
4.6 Variogram model selection
We will use the package gstat and automap for variogram model selection
# specify coordinates in the dataset
coordinates(a) = ~COL+ROW
# select the best model out of exponential, spherical, and gaussian
autofitVariogram(YI ~ COL + ROW, a, model = c("Sph", "Exp", "Gau"), cutoff = 12)## $exp_var
## np dist gamma dir.hor dir.ver id
## 1 264 1.000000 233.2400 0 0 var1
## 2 242 1.414214 388.5222 0 0 var1
## 3 680 2.152750 612.6985 0 0 var1
## 4 812 3.036881 756.8971 0 0 var1
## 5 1066 3.944315 783.1461 0 0 var1
## 6 1364 4.977586 742.6252 0 0 var1
##
## $var_model
## model psill range
## 1 Nug 0.0000 0.000000
## 2 Sph 782.9935 4.019145
##
## $sserr
## [1] 1247749
##
## attr(,"class")
## [1] "autofitVariogram" "list"## np dist gamma dir.hor dir.ver id
## 1 506 1.198102 307.5054 0 0 var1
## 2 680 2.152750 612.6985 0 0 var1
## 3 812 3.036881 756.8971 0 0 var1
## 4 552 3.742751 784.3027 0 0 var1
## 5 834 4.280245 782.1560 0 0 var1
## 6 1044 5.132514 730.3844 0 0 var1
## 7 1028 6.012860 693.9058 0 0 var1
## 8 878 6.801676 675.9157 0 0 var1
## 9 836 7.525735 703.4337 0 0 var1
## 10 852 8.302717 728.0099 0 0 var1
## 11 792 9.194510 712.3311 0 0 var1
## 12 542 10.047104 621.2100 0 0 var1
## 13 452 10.826377 554.3985 0 0 var1
## 14 208 11.494850 599.2237 0 0 var1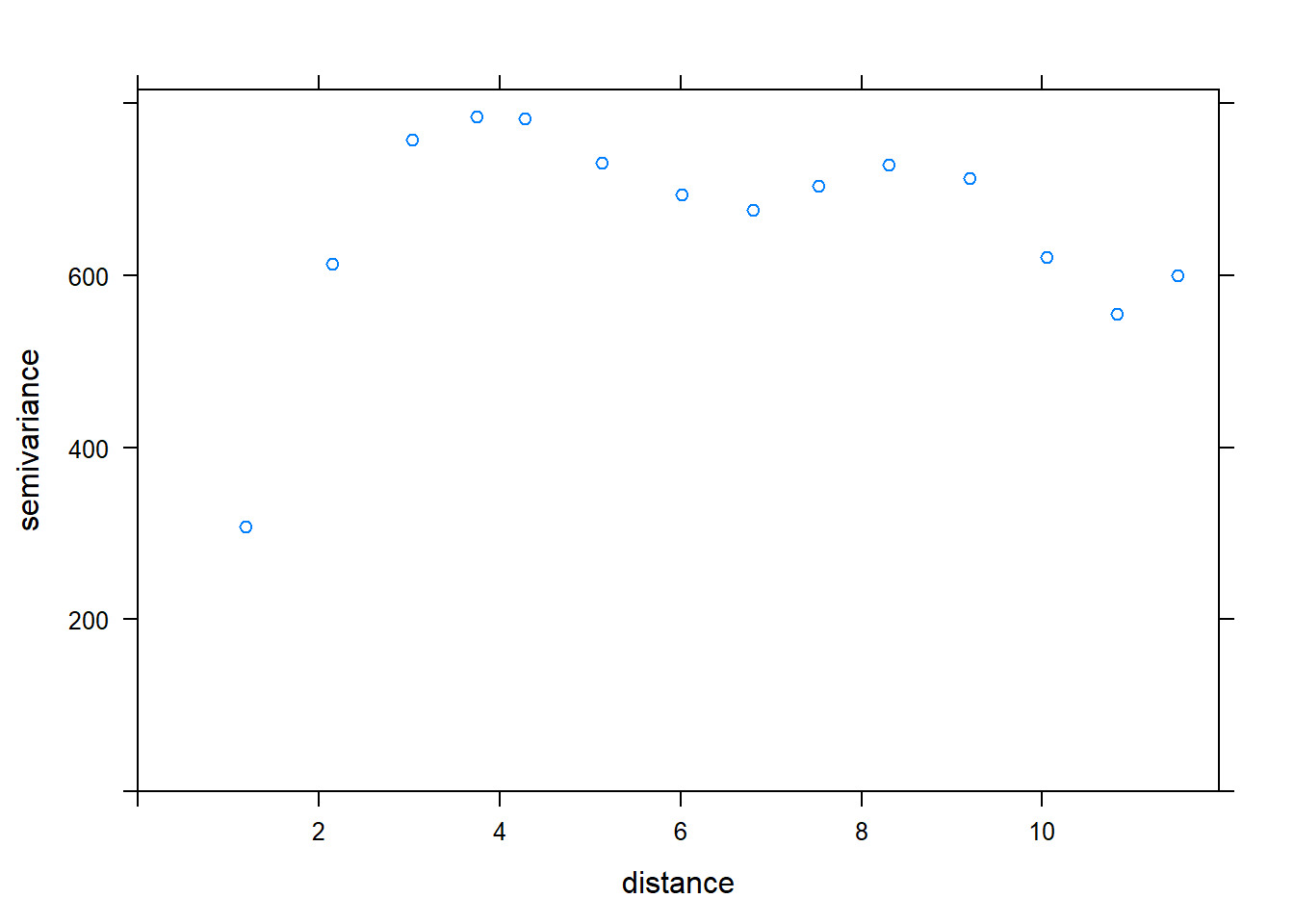
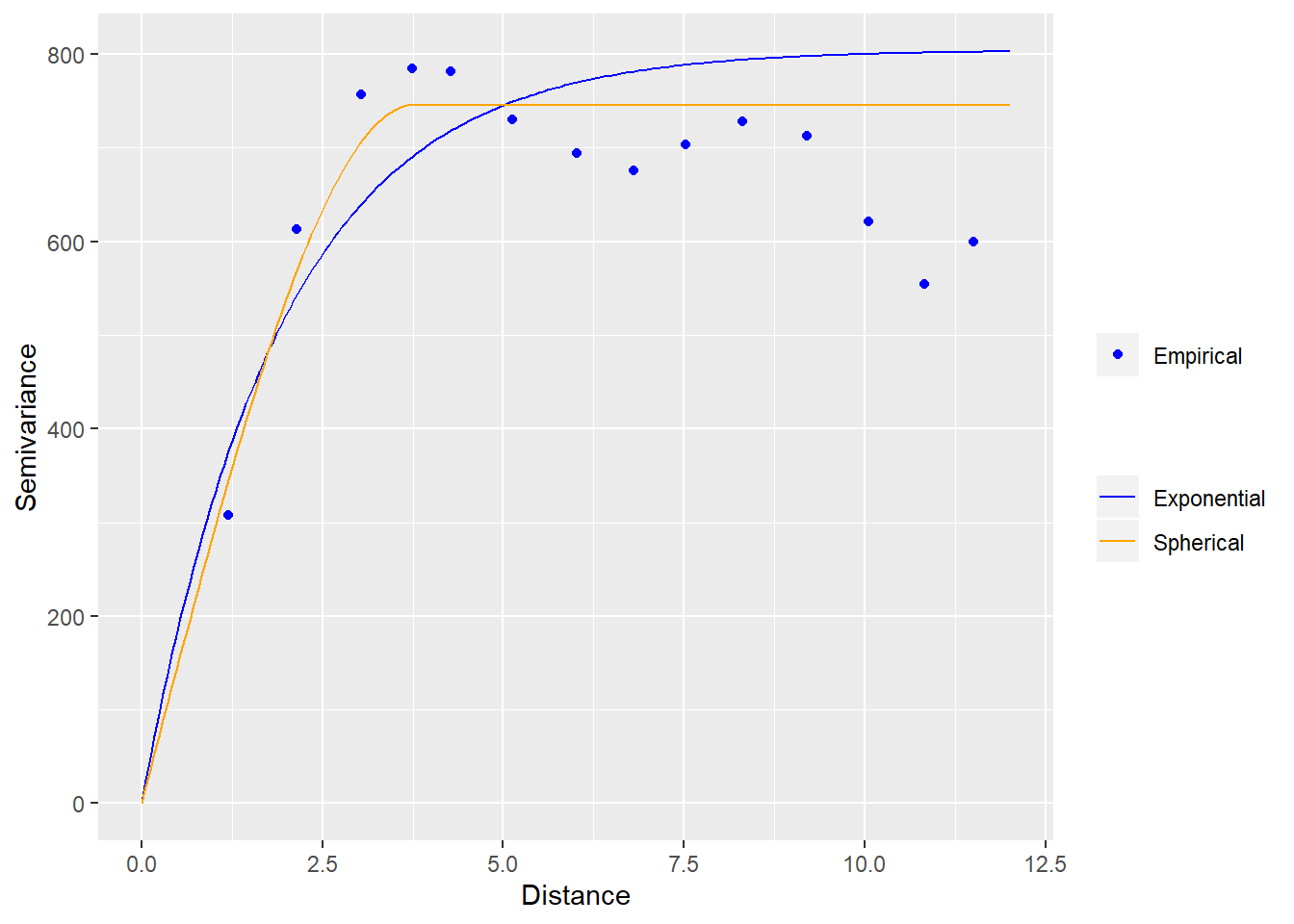
# fit exponential variogram
v_exp <- fit.variogram(v_emp, vgm("Exp"))
# fit spherical and gaussian
v_sph <- fit.variogram(v_emp, vgm("Sph"))
v_sph## model psill range
## 1 Nug 0.0000 0.000000
## 2 Sph 745.8602 3.765221## Warning in fit.variogram(v_emp, vgm("Gau")): No convergence after 200
## iterations: try different initial values?# extract plotting data from fitted variograms
v_exp_line <- variogramLine(v_exp, maxdist = 12)
v_sph_line <- variogramLine(v_sph, maxdist = 12)
# v_gau_line <- variogramLine(v_gau, maxdist = 12)
# plot emprical and fitted variograms together
# specify color for legends
legend_color <- c("Empirical" = "blue", "Exponential" = "blue",
"Spherical" = "orange")
ggplot(data = v_emp) +
geom_point(mapping = aes(x = dist, y = gamma, fill = "Empirical"), color = "blue") +
geom_line(data = v_exp_line, mapping = aes(x = dist, y = gamma, color = "Exponential")) +
geom_line(data = v_sph_line, mapping = aes(x = dist, y = gamma, color = "Spherical")) +
# geom_line(data = v_gau_line, mapping = aes(x = dist, y = gamma, color = "Gaussian")) +
scale_color_manual(name = "", values = legend_color) +
scale_fill_manual(name = "", values = legend_color) +
labs(x = "Distance",
y = "Semivariance")